By Emma Martin, former Analyst at XAnge
AI in biology, drug discovery, or medical applications isn’t new. The subject has been covered extensively in the last years, and models predicting protein binding, estimating drug toxicity, or classifying medical images have already made their way to market. These models have delivered real breakthroughs and efficiency gains.
What’s changing is how these models are built. Instead of building models, that follow a task-by-ask approach, trained to solve one specific, well-defined problem at a time, some are moving toward foundation models. Foundations models are well known in general AI (like GPT or Gemini), but their use in biology has grown rapidly, with more startups and academic papers focusing on them.
To remind the strict definition of foundation models, such as the one stated above, they are large models pre-trained on vast and diverse biological datasets, using self-supervised learning. Instead of learning a single task, they learn general patterns, and relationships across the vast dataset they are fed, and can later be adapted to many downstream applications using relatively little additional data.
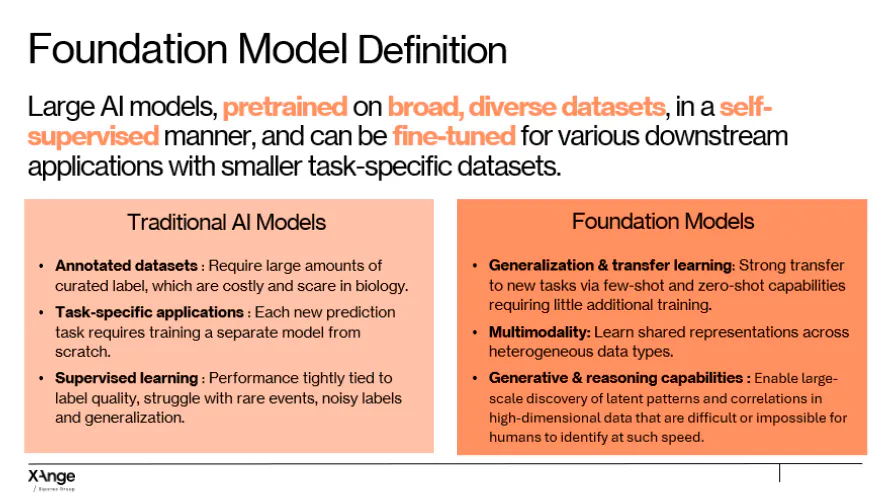

This type of models are particularly useful in biology because biology operates across multiple interconnected levels, from genes and proteins to cells, tissues, and organs, and biological data comes in many forms, including molecular sequences, medical images, and clinical records. Foundation models can integrate these scales and modalities, revealing patterns that span both known and hidden relationships.
Key capabilities of these models include:
- Generalization: They can handle new tasks with few or no examples.
- Multimodality: They can work with different types of data, like molecular sequences, images, and clinical records, all in one model.
- Generative and reasoning power: They can design new molecules, proteins, or antibodies, and simulate biological responses instead of just classifying existing data.
Since 2022, over 200 biological foundation models have been released, alongside major investments. Chai Discovery raised $130M at a $1.3B valuation. Bioptimus secured $76M to build a universal biological foundation model across histology, transcriptomics, and clinical data. Isomorphic Labs, a spin out of Google DeepMind, raised $600M in its first external funding round led by Thrive Capital to bring foundation models further into drug discovery. Capital is concentrating around teams that control unique multimodal datasets, build generalizable models, and demonstrate early commercial traction through pharma partnerships.
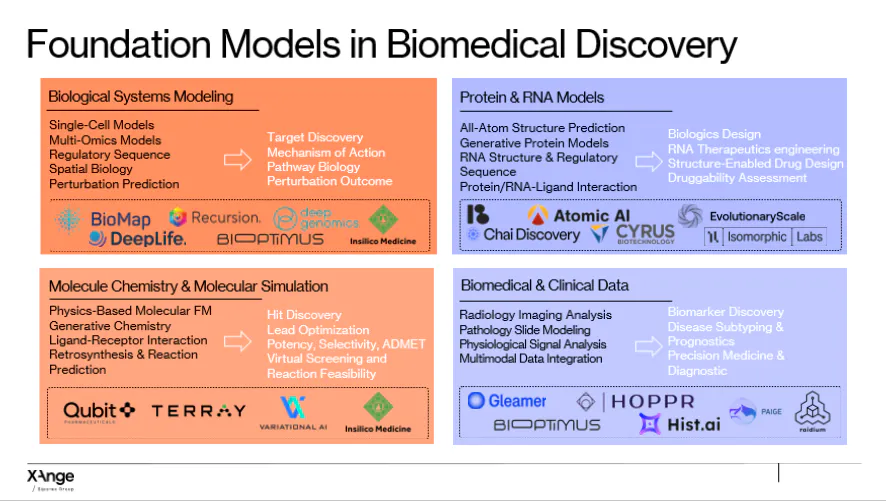
How Bio-Foundation Models Are Used
By Emma Martin, former Analyst at XAnge
Understanding biological systems
These models can predict how genes are expressed, track transitions in cell states, model interactions between immune cells and tumors, and simulate responses to drugs or gene edits. By combining molecular, spatial, and other biological data, they enable experiments in silico that would be difficult or impossible to perform in the lab.
Example: Noetik develops multimodal models of cancer that integrate spatial omics, imaging, and molecular data to simulate tumor–immune dynamics and test hypothetical interventions.
Proteins and RNA
By learning patterns from large datasets of sequences and structures, these models can predict how proteins and RNA fold, interact, and function. They can design antibodies or enzymes, simulate effects of mutations or chemical modifications, and identify binding sites.
Example: Chai Discovery applies zero-shot learning to design high-affinity antibodies without extensive target-specific data. Cradle focuses on generative protein design optimized for both biological function and manufacturability.
Chemistry and drug design
Bio-foundation models can generate novel molecules, predict stability and binding, optimize pharmacological properties, and propose synthetic routes for candidate compounds.
Example: Qubit Pharmaceuticals combines AI with physics-inspired simulations to model molecular interactions for complex or previously intractable targets.
Medical imaging and patient data
These models integrate imaging, molecular, and clinical data to detect subtle biomarkers, predict disease progression, and improve patient stratification for trials.
Example: Owkin’s models, trained on histopathology, omics, and clinical outcomes, help identify predictive biomarkers and guide clinical trial design.

How Companies Are Using Foundation Models
Across Europe, more and more companies are leveraging foundation models, some are building these models as their main product, others are adding them to existing platforms, and some are using them internally to obtain better performance. Looking at companies with public evidence of using foundation models, we can see different ways of leveraging this technology across biomedical research and drug discovery :
- FM-native companies, such as Bioptimus or Raidium, building large multimodal foundation models as their core product, giving them full control over architecture and data and strong long-term defensibility, but at the cost of heavy capital, compute, and long development cycles.
- FM-adopters and specializers, either building foundation models in house for specific applications or integrating pretrained foundation models into existing workflows on specific, domain-focused data to improve performance and speed iteration, benefiting from lower cost and faster deployment but remaining constrained by the limits of the underlying models.
Data as a Moat
The technical architecture of foundation models is increasingly open, what isn't replicable is the data. In biology, the most valuable datasets are built through years of proprietary experiments, clinical partnerships, or exclusive access to patient cohorts, and cannot be matched by a competitor with more compute alone. This is why capital is concentrating around teams that treat data acquisition as a core strategic activity. FM-native companies like Bioptimus are building data flywheels: more partners mean more diverse training data, which improves the model, which attracts more partners. The companies commanding the highest valuations are not just betting on model architecture, they are betting on a compounding data advantage that gets harder to replicate over time.
Why Multimodal Models Matter
Multimodal BioFMs represent a structural shift because these models can jointly learn from molecular data, tissue-level signals and patient-level outcomes, and can capture cross-scale interactions.
We believe this new generation of models has real potential to improve drug development, and specifically in an area that remains a bottleneck: clinical trials. One of the most promising directions is the development of virtual or digital twin models, which aim to simulate biological systems, tissues, or even groups of patients in silico. This matters because most drugs fail late in development because of patient differences, unexpected toxicity, and complex system-level biology. By combining multiple types of data, foundation models make it increasingly realistic to build virtual patient cohorts that can be used to test hypotheses, explore dose–response effects, identify likely non-responders, or surface safety risks before entering expensive Phase II or III trials. These models will not replace clinical trials, but they can help teams fail earlier, cheaper, and with better insight, addressing a part of drug development that earlier AI approaches largely left untouched
Although multimodal biofoundation models are emerging, most models today still focus on a subset of modalities (e.g. omics + images) rather than fully integrating molecular, spatial, clinical, and longitudinal data in a unified way. Training multimodal models requires paired datasets that are rare and difficult to collect. They still rely on scarce, expensive experiments, but on the most optimistic side, the generative ability of these models allow to generate synthetic data, that can partially mitigate this bottleneck by accelerating experiments and amplifying existing data, but this remains an active area rather than a solved problem.
Multimodal Bio-Foundation Models will not eliminate drug discovery failure, but they are the first AI systems structurally capable of addressing its root causes. If they succeed, the impact will not be faster discovery alone, but a fundamental shift in how clinical risk is understood and managed.
By Emma Martin, former Analyst at XAnge
If you want to dive further into this subject, here are some amazing sources that have been great inspirations to write this article :
- On AI Infrastructure in Biology — The Century of Biology
- What are Foundation Models in Biology and Healthcare? — BioPharmaTrend
- AI in Drug Discovery: Use Cases, Market, and Funding in 2025 — Xenoss
- Biological Foundation Models — bioRxiv preprint — bioRxiv
- Why AI Drug Discovery Underperforms in Real Therapeutics and What Needs to Change — Medvolt.ai `
- A New Breed of Biotech — Alex's Blog
- Foundation Models vs. Proprietary Datasets — Where Is the Real Value? — LinkedIn / Yasir Hassan
- Protein Language Models: Builders — Tech x Life Sciences
- Foundational Models Map — Code Ocean
- The Patient is Not a Document: Moving from LLMs to a World Model for Oncology — Standard Model
- Will AI Revolutionize Drug Development? These Are the Root Causes of Drug Failure It Must Address — Singularity Hub
- 2025: The State of AI in Healthcare — Menlo Ventures



